
| Predictor | Mitochondrial | Other | Weight | |
|---|---|---|---|---|
|
Mitoprot
(?)
Score represents the predicted probability that the protein is targeted to mitochondria (mitochondrial score) or not (other score) according to Mitoprot (click to reach). If multiple isofoms of the same protein are available the highest scores are shown for each compartment. Probability values >0.95 are multiplied by 2 as a bonus for the score. If multiple proteins are associated with the reaction, the maximum score for each compartment is presented. See corresponding gene pages (see table below) for the scores of individual proteins. |
2.00 | 1.91 | 2.0 | |
|
MitoMiner
(?)
Score evaluates the evidence for mitochondrial targeting based on the orthologs in 12 eukaryotic model organisms that are in the MitoMiner database (click to reach). Takes a value between 0 and 1 as a function of the ratio of orthologs that have hits (contributes to the mitochondrial score) or no hits (determines the other score) in the database. Mitochondrial score is also a function of the strength of the evidence for the orthologs found in the database (experimental evidence is given a higher weight than theoretical evidence during scoring). If multiple proteins are associated with the reaction, the maximum score for each compartment is presented. See corresponding gene pages (see table below) for the scores of individual proteins. |
1.00 | 0.67 | 1.5 |
Specific protein |
|
UniProt
(?)
Score shows the subcellular localization of proteins encoded by this gene as reported in UniProt (click to reach) in October 2014. Takes a value of either 0 (not reported to be localized to the pertaining compartment) or 1 (reported to be localized to the pertaining compartment). If multiple proteins are associated with the reaction, the maximum score for each compartment is presented. See corresponding gene pages (see table below) for the scores of individual proteins. |
1 | 1 | 1.0 | |
|
Org. DB
(?)
Score shows the subcellular localization of proteins encoded by this gene as reported in Organelle Database (click to reach). Takes a value of either 0 (not reported to be localized to the pertaining compartment) or 1 (reported to be localized to the pertaining compartment). If multiple proteins are associated with the reaction, the maximum score for each compartment is presented. See corresponding gene pages (see table below) for the scores of individual proteins. |
Not available | Not available | 1.0 | |
|
BRENDA
(?)
Score represents the proportion of proteins experimentally localized to the indicated compartment as reported in the BRENDA database (click to reach) for the generic enzyme associated with this reaction. If one of the reported localization experiments for a compartment is carried out in C. elegans, a score of 1.0 is granted regardless of reports from other organisms. If multiple enzymes are associated with the reaction, the maximum score for each compartment is presented. |
0.94 | 0.70 | 1.0 | Generic enzyme |
| BiGG (?) Score represents 0.5 times the number of eukaryotic models with the indicated compartment for this reaction in the BiGG database. Since there are two eukaryotic models, the maximum score is 1.0. | Not available | Not available | 1.5 | Reaction |
|
FBA
(?)
Score shows whether the reaction is useful when it is placed in the indicated compartment based on flux balance analysis. A score of 0 generally indicates no flux carrying capacity, while 1 indicates that the reaction can carry flux. In some cases, a score of 0 also indicates flux carrying as part of a looping pathway, as a looping flux flow does not warrant the contribution of the reaction to metabolic processes in a comparment. If the flux also contributes to biomass or energy generation when placed in the pertaining compartment, a bonus score of 1 is added to reach a maximum score of 2. This score is valid only for PRIME model reactions, and therefore, is "not applicable" to others. |
1 | 0 | 2.0 | |
| Overall (?) Obtained by multiplying each score by the corresponding weight and summing the products. A value >6.2 is considered as sufficient evidence for localization to the pertaining compartment. If neither compartment reaches this threshold, the greater score decides. However, if two scores are within 1.2 of each other, reaction is considered to be localized to both compartments. If FBA score is not available, the first threshold is reduced to 4.2. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 9.4 | 6.5 |
| Source | Forward | Backward | Reversible |
|---|---|---|---|
| MetaCyc (?) Score of 1 represents the direction of the reaction as indicated in MetaCyc. | 0 | 0 | 1 |
| SEED (?) Score of 1 represents the direction of the reaction as indicated in the SEED database. | 0 | 0 | 1 |
| BiGG (?) Score represents 0.5 times the number of models with the indicated direction for this reaction in the BiGG database. | 0.0 | 0.5 | 0.0 |
| BRENDA (?) Score represents the proportion of reactions with the indicated direction reported in the BRENDA database for the enzymes associated with this reaction. Only reactions with a reported direction were used in the calculation. | 0.00 | 0.00 | 1.00 |
| Total (?) If no direction has >0.5 points, reaction is considered reversible. If the score of one direction with the highest score is more than 25% larger than the next highest score, the direction with the highest score is taken. Else, reaction is considered reversible. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 0.00 | 0.50 | 3.00 |
| Source | Forward | Backward | Reversible |
|---|---|---|---|
| MetaCyc (?) Score of 1 represents the direction of the reaction as indicated in MetaCyc. | Not available | Not available | Not available |
| SEED (?) Score of 1 represents the direction of the reaction as indicated in the SEED database. | 0 | 0 | 1 |
| BiGG (?) Score represents 0.5 times the number of models with the indicated direction for this reaction in the BiGG database. | Not available | Not available | Not available |
| BRENDA (?) Score represents the proportion of reactions with the indicated direction reported in the BRENDA database for the enzymes associated with this reaction. Only reactions with a reported direction were used in the calculation. | Not available | Not available | Not available |
| Total (?) If no direction has >0.5 points, reaction is considered reversible. If the score of one direction with the highest score is more than 25% larger than the next highest score, the direction with the highest score is taken. Else, reaction is considered reversible. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 0.00 | 0.00 | 1.00 |
| Source | Forward | Backward | Reversible |
|---|---|---|---|
| MetaCyc (?) Score of 1 represents the direction of the reaction as indicated in MetaCyc. | Not available | Not available | Not available |
| SEED (?) Score of 1 represents the direction of the reaction as indicated in the SEED database. | 0 | 0 | 1 |
| BiGG (?) Score represents 0.5 times the number of models with the indicated direction for this reaction in the BiGG database. | Not available | Not available | Not available |
| BRENDA (?) Score represents the proportion of reactions with the indicated direction reported in the BRENDA database for the enzymes associated with this reaction. Only reactions with a reported direction were used in the calculation. | Not available | Not available | Not available |
| Total (?) If no direction has >0.5 points, reaction is considered reversible. If the score of one direction with the highest score is more than 25% larger than the next highest score, the direction with the highest score is taken. Else, reaction is considered reversible. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 0.00 | 0.00 | 1.00 |
| Source | Forward | Backward | Reversible |
|---|---|---|---|
| MetaCyc (?) Score of 1 represents the direction of the reaction as indicated in MetaCyc. | 0 | 0 | 1 |
| SEED (?) Score of 1 represents the direction of the reaction as indicated in the SEED database. | 0 | 0 | 1 |
| BiGG (?) Score represents 0.5 times the number of models with the indicated direction for this reaction in the BiGG database. | 0.0 | 0.0 | 0.5 |
| BRENDA (?) Score represents the proportion of reactions with the indicated direction reported in the BRENDA database for the enzymes associated with this reaction. Only reactions with a reported direction were used in the calculation. | 0.00 | 0.00 | 1.00 |
| Total (?) If no direction has >0.5 points, reaction is considered reversible. If the score of one direction with the highest score is more than 25% larger than the next highest score, the direction with the highest score is taken. Else, reaction is considered reversible. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 0.00 | 0.00 | 3.50 |
| Source | Forward | Backward | Reversible |
|---|---|---|---|
| MetaCyc (?) Score of 1 represents the direction of the reaction as indicated in MetaCyc. | 0 | 1 | 0 |
| SEED (?) Score of 1 represents the direction of the reaction as indicated in the SEED database. | 0 | 0 | 1 |
| BiGG (?) Score represents 0.5 times the number of models with the indicated direction for this reaction in the BiGG database. | 0.0 | 0.0 | 2.0 |
| BRENDA (?) Score represents the proportion of reactions with the indicated direction reported in the BRENDA database for the enzymes associated with this reaction. Only reactions with a reported direction were used in the calculation. | Not available | Not available | Not available |
| Total (?) If no direction has >0.5 points, reaction is considered reversible. If the score of one direction with the highest score is more than 25% larger than the next highest score, the direction with the highest score is taken. Else, reaction is considered reversible. Exceptions (i.e., cases when these rules are overridden by other criteria) are idicated in general comments to the reaction. | 0.00 | 1.00 | 3.00 |
1.3.8.7Name: medium-chain acyl-CoA dehydrogenase Other names: fatty acyl coenzyme A dehydrogenase (ambiguous), acyl coenzyme A dehydrogenase (ambiguous), acyl dehydrogenase (ambiguous), fatty-acyl-CoA dehydrogenase (ambiguous), acyl CoA dehydrogenase (ambiguous), general acyl CoA dehydrogenase (ambiguous), medium-chain acyl-coenzyme A dehydrogenase, acyl-CoA:(acceptor) 2,3-oxidoreductase (ambiguous), ACADM (gene name). click to see details on this item 5.3.3.8Name: dodecenoyl-CoA isomerase Other names: dodecenoyl-CoA Delta-isomerase, Delta3-cis-Delta2-trans-enoyl-CoA isomerase, acetylene-allene isomerase, dodecenoyl-CoA Delta-isomerase, dodecenoyl-CoA Delta3-cis-Delta2-trans-isomerase click to see details on this item 4.2.1.17Name: enoyl-CoA hydratase Other names: enoyl hydrase, unsaturated acyl-CoA hydratase, beta-hydroxyacyl-CoA dehydrase, beta-hydroxyacid dehydrase, hydratase, enoyl coenzyme A, acyl coenzyme A hydrase, crotonase, crotonyl hydrase, 2-octenoyl coenzyme A hydrase, enoyl coenzyme A hydratase, 2-enoyl-CoA hydratase, short-chain enoyl-CoA hydratase, ECH, trans-2-enoyl-CoA hydratase, enoyl coenzyme A hydrase (D), enoyl coenzyme A hydrase (L), short chain enoyl coenzyme A hydratase, D-3-hydroxyacyl-CoA dehydratase, enol-CoA hydratase click to see details on this item 1.1.1.211Name: long-chain-3-hydroxyacyl-CoA dehydrogenase Other names: beta-hydroxyacyl-CoA dehydrogenase, long-chain 3-hydroxyacyl coenzyme A dehydrogenase, 3-hydroxyacyl-CoA dehydrogenase, LCHAD click to see details on this item 1.1.1.35Name: 3-hydroxyacyl-CoA dehydrogenase Other names: beta-hydroxyacyl dehydrogenase, beta-keto-reductase, 3-keto reductase, 3-hydroxyacyl coenzyme A dehydrogenase, beta-hydroxyacyl-coenzyme A synthetase, beta-hydroxyacylcoenzyme A dehydrogenase, beta-hydroxybutyrylcoenzyme A dehydrogenase, 3-hydroxyacetyl-coenzyme A dehydrogenase, L-3-hydroxyacyl coenzyme A dehydrogenase, L-3-hydroxyacyl CoA dehydrogenase, beta-hydroxyacyl CoA dehydrogenase, 3beta-hydroxyacyl coenzyme A dehydrogenase, 3-hydroxybutyryl-CoA dehydrogenase, beta-ketoacyl-CoA reductase, beta-hydroxy acid dehydrogenase, 3-L-hydroxyacyl-CoA dehydrogenase, 3-hydroxyisobutyryl-CoA dehydrogenase, 1-specific DPN-linked beta-hydroxybutyric dehydrogenase click to see details on this item 2.3.1.16Name: acetyl-CoA C-acyltransferase Other names: beta-ketothiolase, 3-ketoacyl-CoA thiolase, KAT, beta-ketoacyl coenzyme A thiolase, beta-ketoacyl-CoA thiolase, beta-ketoadipyl coenzyme A thiolase, beta-ketoadipyl-CoA thiolase, 3-ketoacyl CoA thiolase, 3-ketoacyl coenzyme A thiolase, 3-ketoacyl thiolase, 3-ketothiolase, 3-oxoacyl-CoA thiolase, 3-oxoacyl-coenzyme A thiolase, 6-oxoacyl-CoA thiolase, acetoacetyl-CoA beta-ketothiolase, acetyl-CoA acyltransferase, ketoacyl-CoA acyltransferase, ketoacyl-coenzyme A thiolase, long-chain 3-oxoacyl-CoA thiolase, oxoacyl-coenzyme A thiolase, pro-3-ketoacyl-CoA thiolase, thiolase I, 2-methylacetoacetyl-CoA thiolase [misleading] click to see details on this item |
[m] : (2) coaName: Coenzyme A Formula: C21H32N7O16P3S Charge: -4 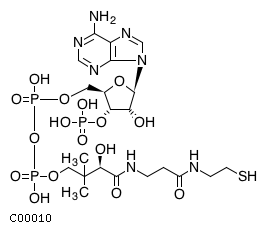 click to see details on this item + (2) h2oName: H2O Formula: H2O Charge: 0 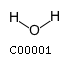 click to see details on this item + (2) nadName: Nicotinamide adenine dinucleotide Formula: C21H26N7O14P2 Charge: -1 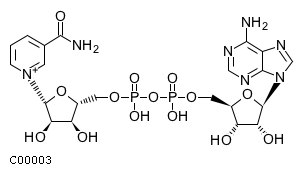 click to see details on this item + fadName: Flavin adenine dinucleotide oxidized Formula: C27H31N9O15P2 Charge: -2 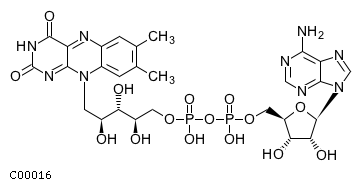 click to see details on this item + arachdcoaName: C20:4-CoA Formula: C41H62N7O17P3S Charge: -4  click to see details on this item |
 |
(2) accoaName: Acetyl-CoA Formula: C23H34N7O17P3S Charge: -4 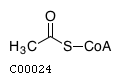 click to see details on this item + (2) hName: H+ Formula: H Charge: 1  click to see details on this item + fadh2Name: Flavin adenine dinucleotide reduced Formula: C27H33N9O15P2 Charge: -2 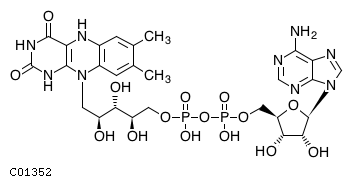 click to see details on this item + (2) nadhName: Nicotinamide adenine dinucleotide - reduced Formula: C21H27N7O14P2 Charge: -2 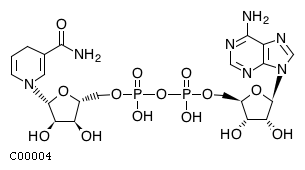 click to see details on this item + fa16p3n6coaName: Intermediary fatty acid C16:3 Formula: C37H56N7O17P3S Charge: -4 click to see details on this item |