
| Abbreviation ..................................................... | : | pheme |
| Name ..................................................... | : | Protoheme, etc.Heme, Haem, Heme B, Protoheme IX |
| Model ID ..................................................... | : | C00032 |
| KEGG ID (?) Compounds in KEGG database that match this compound. ..................................................... | : |
C00032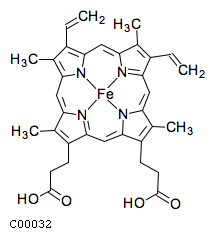
|
| Formula ..................................................... | : | C34H30FeN4O4 |
| Charge ..................................................... | : | 0 |
| Localization ..................................................... | : | Mitochondria, Cytosol, Extracellular |
| BiGG matches (?) Compounds in BiGG database that match this compound. ..................................................... | : | pheme |
| Other associations ..................................................... | : | See reaction table below (8 reactions in total) |
|
|
||
|
In WormPaths
(?)
Pathway maps where this metabolite is represented are listed here. Letters after map name indicate the compartments the metabolite is placed in the map: (c) Cytosol (m) Mitochondria (e) Extracellular space ..................................................... |
: | Iron metabolism (c,e,m) |